How to Perform Lasso Regression in R
The Lasso Regression is a classification algorithm that uses the sparse and shrinkage in simple models.
This tutorial demonstrates how to perform lasso regression in R.
Lasso Regression in R
The LASSO is abbreviated as Least Absolute Shrinkage and Selection Operator. When we want to automate certain parts of model selection, the lasso regression is a good choice as it shows a high level of multicollinearity.
The lasso regression is a quadratic programming problem, and the languages like R and Matlab provide solutions. Let’s see step by step process to solve lasso regression in R.
Understanding the Equation
The Lasso regression minimizes the following function.
RSS + λΣ|βj|
Where j is the range from 1 to the predictor variable and the λ ≥ 0, the second term λΣ|βj| is known as shrinkage penalty.
The RSS = Σ(Yi – ŷi)2, in which Σ is the sum, yi is the actual response value for ith observation, and ŷi is the predicted response value.
The lambda is selected as the lowest possible test mean squad error (MSE) in lasso regression once we know what we will do in the lasso regression.
Let’s load the data in the next step.
Load the Data
Let’s use the mtcars dataset for our example. The hp will be used as the response variable and mpg, drat, wt, qsec as the predictors.
We can use the glmnet package to perform the lasso regression. Let’s load the data.
# glmnet package requires to define response variable
x <- mtcars$hp
# glmnet package requires to define matrix of predictor variables
y <- data.matrix(mtcars[, c('mpg', 'wt', 'drat', 'qsec')])
Once data is loaded, the next is to fit the lasso regression model.
The Lasso Regression Model Fitting
In this step, we use the glmnet() function to fit the lasso regression model; the alpha will be set to 1 for the lasso regression model. The k-fold cross-validation will be performed to determine the value of lambda, and with glmnet, it automatically performs cross-validation with k= 10 folds.
We might need to install the glmnet package if it is already not installed. See example:
# k-fold cross-validation to find lambda value
cv_model <- cv.glmnet(x, y, alpha = 1)
#lambda value that minimizes test MSE
best_lambda <- cv_model$lambda.min
best_lambda
# plot the model
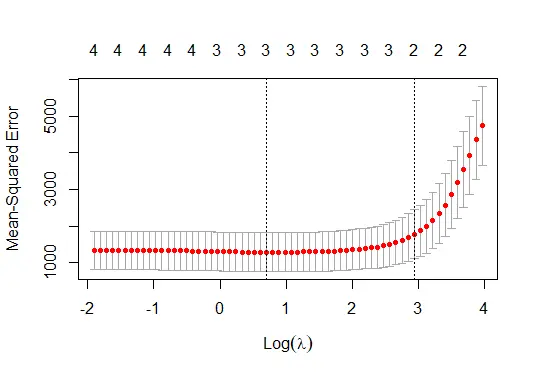
plot(cv_model)
The code above fits the lasso regression model and shows the best lambda value and the model plot.
The lambda value which minimizes the test MSE is below.
[1] 2.01841
The lasso regression model plot:
Analyze the Lasso Regression Model
Analyzing the model means we can show to coefficients of the model. The coefficients for our model are:
# Coefficients of the model
coef(cv_model)
Output for the code will be:
5 x 1 sparse Matrix of class "dgCMatrix"
s1
(Intercept) 418.277928
mpg -4.379633
wt .
drat .
qsec -10.286483
We know the best lambda value; we can also create the best model bypassing the best lambda value as the argument to the glmnet function while fitting the model.
As we can see, no coefficients are shown for the predictors wt and drat because the lasso regression shrunk them to 0. That is why they were dropped from the model.
Sheeraz is a Doctorate fellow in Computer Science at Northwestern Polytechnical University, Xian, China. He has 7 years of Software Development experience in AI, Web, Database, and Desktop technologies. He writes tutorials in Java, PHP, Python, GoLang, R, etc., to help beginners learn the field of Computer Science.
LinkedIn Facebook